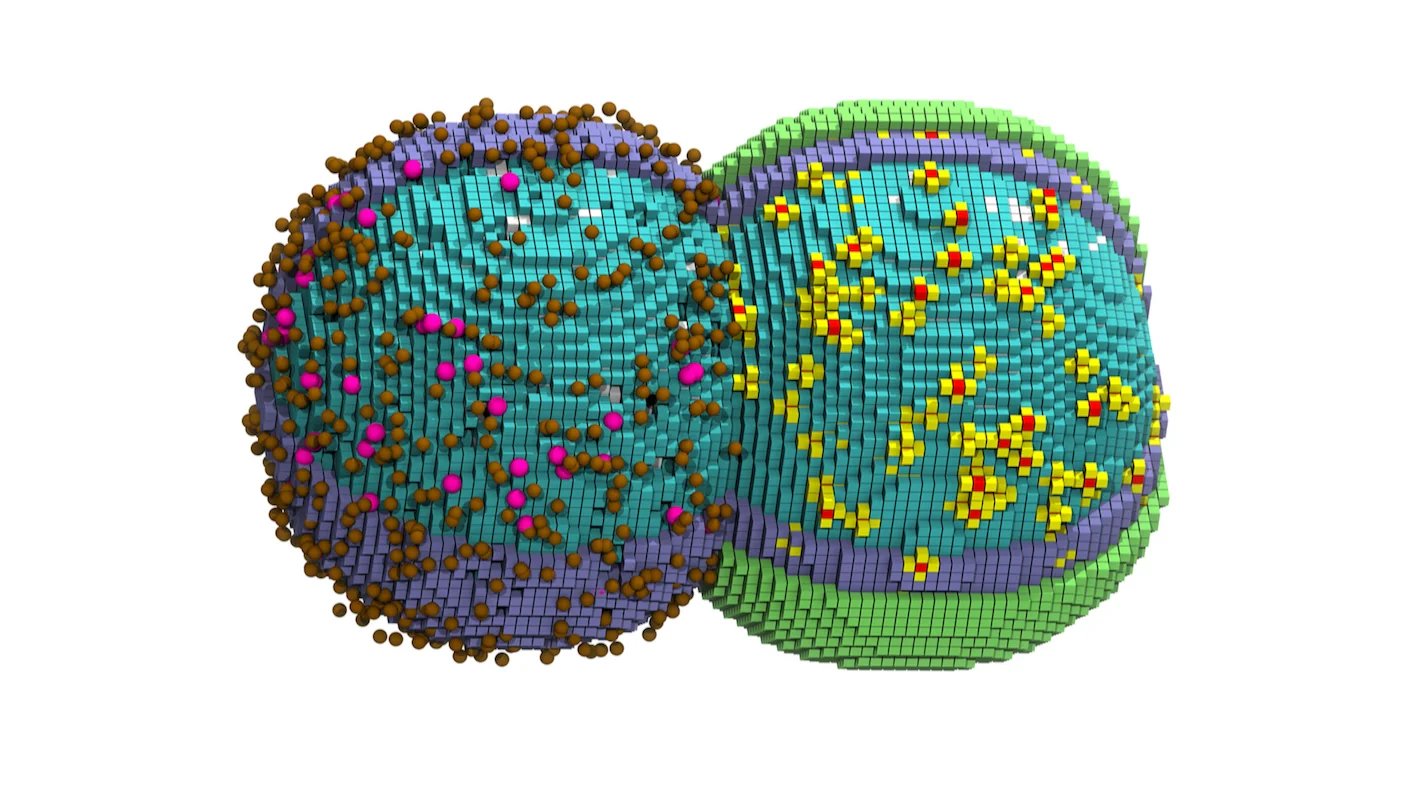
Five years ago, scientists watched in wonder as synthetic bacteria grew and split into daughter cells. The bacteria’s extremely stripped-down genome still supported its entire life cycle. It was a crowning achievement in synthetic biology that shed light on life’s most basic processes.
These processes can now be viewed digitally. This month, a team at the University of Illinois at Urbana-Champaign developed a virtual model of the bacteria tracking nearly all of a cell’s molecules down to the nanoscale. The researchers made this digital cell by combining several large datasets covering thousands of molecules and then animating them as the bacteria split in two.
The model is the latest in a growing effort to make digital twins of living cells. Mimicking diseases or treatments in the digital world offers a bird’s-eye view of cellular changes and could speed up drug discovery and help researchers tackle complex diseases like cancer.
“We have a whole-cell model that predicts many cellular properties simultaneously,” study author Zan Luthey-Schulten said in a press release. The model could provide “the results of hundreds of experiments” at the same time, she said.
Digitizing Life
Every cell is a bustling metropolis. Proteins orchestrate a vast range of cellular responses. RNA molecules carry instructions from genes to the cell’s protein-building factories. Fatty acids in a cell’s membrane rearrange themselves to admit nutrients or ward off invaders. Working in tandem, they all keep the cell humming along.
This complexity makes cells hard to simulate. But with large datasets charting the genome, gene expression, and proteins alongside sophisticated AI, scientists have built static virtual cells that paint a near-complete picture with atomic-level resolution. More recent models can even predict molecular movements for a short period of time (often less than a second).
But they can’t simulate “the mechanics and chemistry that take place over minutes to hours in processes such as gene expression and cell division,” wrote the University of Illinois team.
Other efforts use physics to predict how molecular changes affect behavior in bacteria, yeast, and human cells. These treat cells as a “well-stirred system”—that is, a cup of molecular soup lacking details about where each molecule sits and how molecules vary from cell to cell.
But location is key. As cells divide, some proteins gather around DNA to help copy it; others assemble near the membrane to recruit fatty molecules for its growth as the cell splits in two.
Simulating everything, everywhere, all at once during human cell division is beyond even the most powerful supercomputers. Minimal bacteria offer an alternative. These synthetic bacteria are stripped-down versions of the parasite Mycoplasma mycoides. The team focused on one of these known as JCVI-syn3A. Its 493-gene genome—roughly half the original—is the smallest set of DNA instructions to boot up a living bacteria that can still grow and divide.
In 2022, the team developed a 3D model of the bacteria’s metabolism, genes, and growth. But the software, Lattice Microbes, struggled to track division.
Life in 4D
The new study added more data to the software. This included membrane changes and information about how ribosomes, the cell’s protein-making machines, assemble and move inside the cell’s gooey interior. They also added stochasticity, or unpredictability, to the model.
Changes to the location of chromosomes, which house DNA, are random as the cell divides, which makes them difficult to predict. But their position influences DNA replication and gene expression.
The first update nearly broke the software. It could map molecules involved in cell division, such as an enzyme critical for DNA copying. But adding chromosome location predictions slowed the model to a crawl, even when running on advanced GPUs. Most of the cells died before their simulations were complete.
Several tweaks helped. One was to add more computational power. The team used a GPU dedicated to chromosomes, while all other details were processed on a separate chip. The model also ran faster by rendering some proteins as inert spheres that could be largely ignored.
The upgrades worked. Leaving the model running over Thanksgiving, the team returned to find it had completed the bacteria’s whole life cycle. “All of a sudden, it was just this huge leap,” study author Zane Thornburg told Nature.
The simulation matched many real-world experiments, such as how the cells elongate and bubble into dumbbell-like shapes during division. The model also accurately predicted the length of a cell cycle and captured a wide range of cellular activity.
“I can’t overstate how hard it is to simulate things that are moving—and doing it in 3D for an entire cell was…triumphant,” said Thornburg.
Every cell is like a snowflake: Although containing similar molecules, the amounts and locations differ. The model easily handled this diversity. Repeated simulations of the bacteria, each starting with slightly different genetic, molecular, and metabolic makeup, resulted in a similar cycle length and movement of chromosomes during division.
The results came at a cost: Simulating the cell’s 105-minute cycle took up to six days on a supercomputer. But the virtual cell could lend insights into the molecular dance that causes all cells to grow and divide. JCVI-syn3A doesn’t have the smallest genome. Its predecessor holds the record, but it also struggles to make normally shaped and functional daughter cells—suggesting some genes are essential for division. Simulation could help us understand why.
Other efforts using generative AI to build virtual cells are in the works. But because this study’s model was grounded in strict physical and biochemical rules, results could be easily verified in the lab. AI-generated virtual cells, however, are commonly trained on gene expression data alone, which is a snapshot of a cell’s state and often fails to predict complex cell responses.
The two approaches could inspire each other by homing in on principles that make a virtual cell run like the real deal. For example, they could show that capturing each molecule in space and time, rather than as a soup, vastly improves the model.
Although the model can’t simulate a cell atom-by-atom, the team wrote, it could “illuminate the interwoven nature of the biology, chemistry, and physics that govern life for cells.”