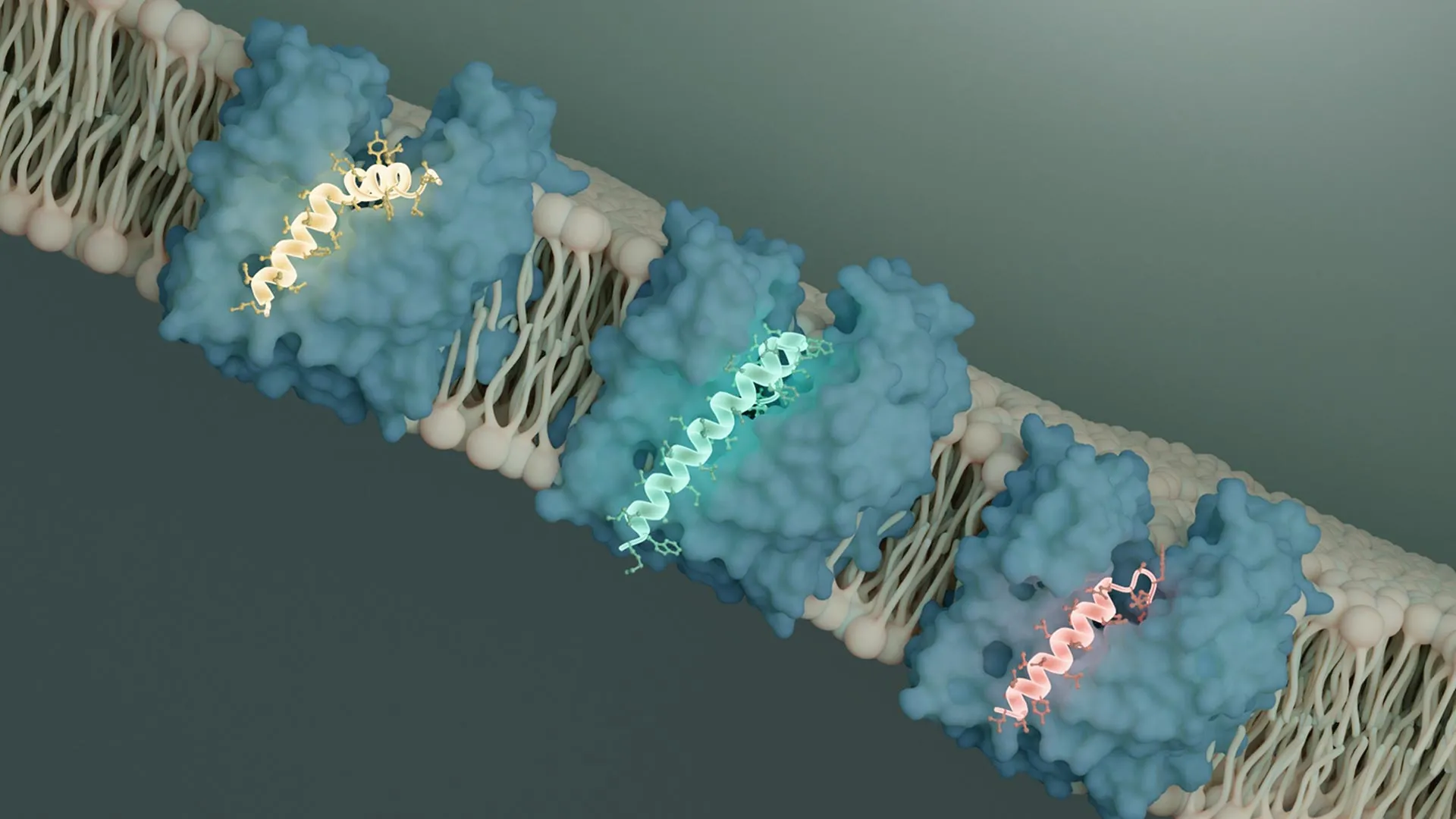
Scientists have revealed how viruses that infect bacteria shut down MurJ, a protein essential for building the bacterial cell wall. Remarkably, different viruses evolved separate proteins that all block MurJ in the same way, highlighting it as a promising new antibiotic target.
The findings appear in the February 26 issue of Nature. The research was led by Yancheng Evelyn Li, a graduate student in the lab of Bil Clemons at Caltech. Clemons, the Arthur and Marian Hanisch Memorial Professor of Biochemistry, is the corresponding author.
The Urgent Need for New Antibiotics
Bacteria evolve quickly, and that adaptability is fueling a growing public health crisis. As Clemons explains, “Evolution is powerful, and in bacteria, resistance to antibiotics develops quickly. This means that we now deal with bacteria that are resistant to all the medicines that we have.” He adds, “In the USA alone, tens of thousands of people die every year from antibiotic-resistant bacterial infections, and that number is rising rapidly. We need new antibiotics to combat this.”
With existing drugs losing effectiveness, researchers are searching for entirely new bacterial weak points.
Targeting the Bacterial Cell Wall
One long standing focus has been the pathway bacteria use to construct peptidoglycan, the rigid material that forms their cell wall. This process, called the peptidoglycan biosynthesis pathway, is especially attractive because peptidoglycan is found in bacteria but not in human cells. As Clemons notes, “Peptidoglycan is a unique feature of bacteria, and that makes it an attractive antibiotic target.”
Several antibiotics already disrupt this pathway. Penicillin, discovered by Alexander Fleming in the mid 20th century, blocks a late stage of peptidoglycan production. Related drugs such as amoxicillin work in a similar way.
Key Proteins MraY, MurG, and MurJ
Three essential proteins drive the movement of peptidoglycan building blocks across the bacterial inner membrane: MraY, MurG, and MurJ. These proteins help transport the components needed to assemble the cell wall outside the inner membrane barrier. If any one of them fails, peptidoglycan cannot be produced and the bacterium dies, making them promising drug targets.
Although researchers understand much about how these proteins function, Clemons points out that important mechanistic details remain unclear.
At present, no approved drugs directly inhibit these three proteins. Still, Clemons says there is potential. “We do know that we can find small molecules, either derived from nature or synthesized in chemical libraries, that will inhibit these proteins. Excitingly, recent discoveries have shown that bacteriophages have figured out how to target this pathway.”
How Bacteriophages Break Through Bacterial Defenses
Bacteriophages, or phages, are viruses that infect bacteria. To survive, they must enter a bacterial cell, replicate, and then escape to infect others. Breaking out requires getting through the peptidoglycan layer. Clemons explains, “Getting back out means that they have to get past the peptidoglycan layer. Because it acts like chainmail, the phages get stuck if they can’t break through it.”
The Clemons lab studies small phages that contain single stranded DNA or RNA. These viruses have compact genomes and rely on simple strategies to kill bacteria. In 2023, the team reported in Science on φX174, a phage with a long research history at Caltech.
Viral Proteins That Disable MurJ
Small phages rely on specialized protein antibiotics called single-gene lysis proteins, or Sgls (pronounced like “sigils”), to kill bacteria. Li and Clemons have focused on Sgls that target MurJ, one of the key cell wall proteins.
MurJ acts as a flippase. It transports peptidoglycan precursors from the inside of the cell across the membrane so they can be incorporated into the growing cell wall. Earlier work from collaborators showed that two unrelated Sgls, SglM and SglPP7, both kill bacteria by blocking MurJ.
To understand how this happens, Li used cryo electron microscopy at Caltech’s Beckman Institute Biological and Cryogenic Transmission Electron Microscopy (Cryo-EM) Resource Center. Flippases such as MurJ move molecules by alternately exposing them to each side of the membrane without forming a permanent opening. When MurJ binds its cargo inside the cell, it changes shape to release the molecule on the outside.
Li found that both SglM and SglPP7 attach to a groove in MurJ, preventing the structural shift required for transport.
“It is clear that both of these Sgls bind to MurJ in an outward-facing conformation, locking it into this position,” Li says. Researchers are encouraged by this because the outward-facing form of MurJ is exposed to the surrounding environment, which could make it more accessible to future drugs than a conformation that faces inward.
Convergent Evolution Highlights a Promising Drug Target
Clemons says the result was surprising for another reason. “These peptides, which have no evolutionary links to each other, have both figured out how to target MurJ in a very similar way. These are two examples of convergent evolution, in which different evolutionary paths arrive at the same solution. We were surprised!”
Because viruses evolve rapidly, the team believes many more phages likely carry similar Sgls. Phages are relatively easy to isolate, and studying their genomes could reveal additional biological insights and new antibiotic targets.
In the Nature study, the researchers analyzed another phage genome with the help of a collaborator. They identified a new Sgl called SglCJ3 (from a genome sequence that is predicted to be a phage and is called Changjiang3) and examined it using cryo electron microscopy. Li determined the structure of SglCJ3 bound to MurJ and found that it also locks the protein in the same outward-facing conformation.
“This is a third genome that evolved a distinct peptide to inhibit the same target in a similar way,” Clemons says. “It is the first strong evidence that evolution identifies MurJ as a great target for killing bacteria, which means we should follow evolution’s lead and develop therapeutics that target MurJ. This demonstrates the power of basic biology to help us solve problems in medicine. Our path is set on leveraging Sgl discovery, and we hope to continue to be supported to turn these concepts into realities.”
Authors and Funding
The paper is titled “Convergent MurJ flippase inhibition by phage lysis proteins.” In addition to Clemons and Li, the authors include Caltech graduate student Grace F. Baron and Francesca S. Antillon, Karthik Chamakura, and Ry Young of Texas A&M University. The research was supported by the Chan Zuckerberg Initiative, the National Institutes of Health, the G. Harold and Leila Y. Mathers Foundation, and the Center for Phage Technology at Texas A&M, jointly sponsored by Texas A&M AgriLife.







![‘Daredevil: Born Again’ Season 2 Finale Explained: [Spoiler]’s Cameo and Predictions for the MCU](https://dailynewsnblog.com/wp-content/uploads/2026/05/marvel-daredevil-born-again-s2-1.jpg)
